Analysis¶
The Analysis class is the primary class which will provide methods for analyzing your life data. This class is designed to take your data and calculate \(\beta\) and \(\eta\) values along with generating any appropriate plots for display of your data.
A typical use case of the Analysis case is as follows:
import weibull
# fail times include no censored data
fail_times = [
9402.7,
6082.4,
13367.2,
10644.6,
8632.0,
3043.4,
12860.2,
1034.5,
2550.9,
3637.1
]
# this is where the actual analysis and curve fitting occur
analysis = weibull.Analysis(fail_times, unit='hour')
Fitting¶
The fit() method is used to calculate appropriate \(\beta\) and \(\eta\) values, which are then stored into the class instance. When fit() is called with no parameters, then the linear regression method of calculation is assumed:
analysis.fit()
An alternative method is to use the Maximum Likelihood Estimation (MLE) method of fitting \(\beta\) and \(\eta\) to the data. This may be done by specifying that the method='mle':
analysis.fit(method='mle')
In many cases, the mle and lr methods will yield very similar values for \(\beta\) and \(\eta\), but there are some cases in which one is preferred over the other. It turns out that linear regression tends to work best for very low sample sizes, usually less than 15 while the maximum likelihood estimator works best for high sample sizes. In both cases, the probplot() method should be used to verify that the data is a good fit.
To retrieve the \(\beta\) and \(\eta\) values, simply use the instance variables beta and eta:
print(f'beta: {analysis.beta: .02f}')
print(f'eta: {analysis.eta: .02f}')
Use the stats() method to get a list of most internal estimates:
$> analysis.stats
fit method maximum likelihood estimation
confidence 0.6
beta lower limit 2.42828
beta nominal 2.97444
beta upper limit 3.64344
eta lower limit 186.483
eta nominal 203.295
eta upper limit 221.622
mean life 181.47
median life 179.727
b10 life 95.401
dtype: object
Plotting¶
One of the most often requested features of such a package is plotting the data, particularly in Jupyter Notebooks. The weibull package comes with built-in methods to easily display and save standard plots with one-line methods.
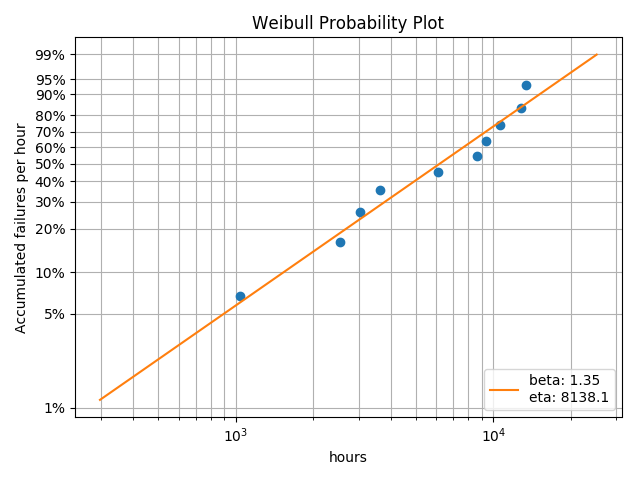
Building on the analysis instance above, we will examine the probability plot:
analysis.probplot()
We can also examine a number of other common function plots (only the hazard plot is shown, but the others are along the same line).:
analysis.pdf()
analysis.sf()
analysis.hazard()
analysis.cdf()
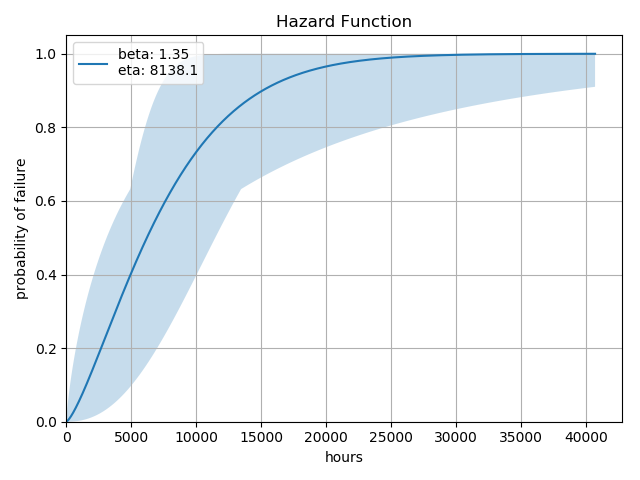
Each of these functions will generate a plot that is suitable for publication or insertion into a Jupyter Notebook. Again, note that some of these methods - such as hazard() and cdf() will produce the same plot with slightly different labeling.
Class Documentation¶
-
class
weibull.Analysis(data: list, suspended: list = None, unit: str = 'cycle')¶ Calculates and plots data points and curves for a standard 2-parameter Weibull for analyzing life data.
Parameters: - data – A list or numpy array of life data, i.e.
[127, 234, 329, 444] - suspended – A list or numpy array of suspensions as boolean values, i.e.
[False, False, True, True]. At any point which indicatesTruemeans that the test was stopped - or that the item was removed from the test - before the item failed. - unit – The unit (‘hour’, ‘minute’, ‘cycle’, etc.). This is used to add some useful information to the visualizations. For instance, if the unit is
hour, then the x-axis will be labed in hours.
Variables: - beta – The current value of the shape parameter, \(\beta\). This value is initially set to
None. The proper value forbetawill be calculated on call to thefit()method. The user may also set this value directly. - eta – The current value of the scale parameter, \(\eta\). This value is initially set to
None. The proper value forbetawill be calculated on call to thefit()method. The user may also set this value directly. - _fit_test – Basic statistics regarding the results of
fit(), such as \(R^2\) and P-value.
-
b(percent_failed: (<class 'float'>, <class 'str'>) = 10.0)¶ Calculate the B-life value
Parameters: percent_failed – the number of elements that have failed as a percent (i.e. 10) Returns: the life in cycles/hours/etc.
-
cdf(show: bool = True, file_name: str = None)¶ Plot the cumulative distribution function
Parameters: - show – True if the plot is to be shown, false if otherwise
- file_name – the file name to be passed to
matplotlib.pyplot.savefig
Returns: None
-
characteristic_life¶ Returns the current characteristic life of the product, aka \(\eta\)
Returns: the characteristic life of the product
-
fit(method: str = 'lr', confidence_level: float = 0.95)¶ Calculate \(\beta\) and \(\eta\) using a linear regression or using the maximum likelihood method, depending on the ‘method’ value.
Method: ‘lr’ for linear estimation or ‘mle’ for maximum likelihood estimation Returns: None
-
fr(show: bool = True, file_name: str = None)¶ Plot failure rate as a function of cycles
Parameters: - show – True if the item is to be shown now, False if other elements to be added later
- file_name – if file_name is stated, then the probplot will be saved as a PNG
Returns: None
-
hazard(show: bool = True, file_name: str = None)¶ Plot the hazard (CDF) function
Parameters: - show – True if the plot is to be shown, false if otherwise
- file_name – the file name to be passed to
matplotlib.pyplot.savefig
Returns: None
-
mean¶ Calculates and returns mean life (aka, the MTTF) is the integral of the reliability function between 0 and inf,
\[MTTF = \eta \Gamma(\frac{1}{\beta} + 1)\]where gamma function, \(\Gamma\), is evaluated at \(\frac{1}{\beta+1}\)
Returns: the mean life of the product
-
median¶ Calculates and returns median life of the product
Returns: The median life
-
mttf¶ Calculates and returns mean time between failures (MTTF)
Returns: the mean time to failure
-
pdf(show: bool = True, file_name: str = None)¶ Plot the probability density function
Parameters: - show – True if the plot is to be shown, false if otherwise
- file_name – the file name to be passed to
matplotlib.pyplot.savefig
Returns: None
-
probplot(show: bool = True, file_name: str = None, **kwargs)¶ Generate a probability plot. Use this to show the data points plotted with the beta and eta values.
Parameters: - show – True if the plot is to be shown, false if otherwise
- file_name – the file name to be passed to
matplotlib.pyplot.savefig - kwargs – valid matplotlib options
Returns: None
-
sf(show: bool = True, file_name: str = None)¶ Plot the survival function
Parameters: - show – True if the plot is to be shown, false if otherwise
- file_name – the file name to be passed to
matplotlib.pyplot.savefig
Returns: None
-
stats¶ Returns the fit statistics, confidence limits, etc :return: a pandas series containing the fit statistics
- data – A list or numpy array of life data, i.e.